An exciting workshop, organised as the capstone of the International Year of Statistics Statistics2013, was celebrated on the 11th and 12th of November at the Royal Statistical Society Headquarters in London, under the stimulating title of “The Future of the Statistical Science”, science to which John Pullinger, president of the Royal Statistical Society, refers to as a quintessentially, multidisciplinary discipline. The workshop was indeed a good example of that.
Online attendance was welcomed to the event giving an additional opportunity to listen to highly reputed professionals in main areas of Statistics. Virtual participants were also allowed to pose their questions to the speakers, an innovation that worked very well as an excellent way to make these sorts of events available to a wider audience that would otherwise be excluded – I am already looking forward to more! -.
In case you missed it at the time, luckily the podcast is still available here.
True to its description, the event covered different areas of application, it showed the tools of the field for tackling a wide range of challenges, it portrayed potentially high impact examples of research, and in summary, was a great attention-grabbing exercise that will hopefully encourage other professionals to the area. A common vision of ubiquity and great relevance came across from the speeches and showed the field as a very attractive world to join. See my simplified summary below in form of a diagram certainly reflective of the great Statistics Views motto “Bringing Statistics Together”.
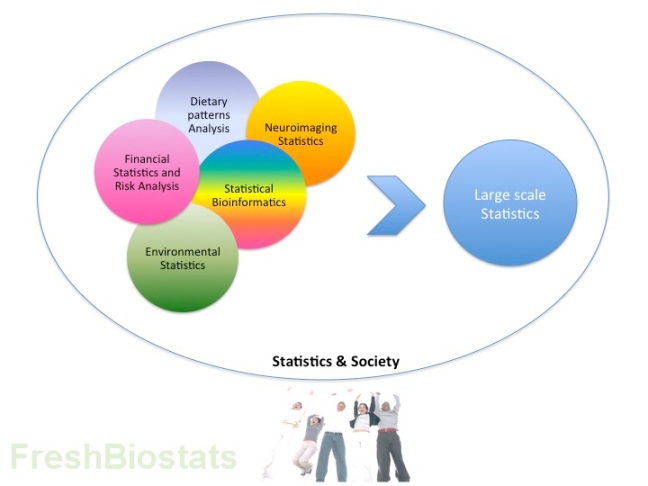
Fig 1. Some of the ideas discussed in the workshop
Particularly relevant to this blog and my particular interests were presentations on environmental Statistics and statistical Bioinformatics, as well as other health-related talks. Important lessons were taken on board from the rest of the speakers too.
The workshop started with a shared presentation on the hot topic “Statistical Bioinformatics” by Peter Bühlmann (ETH Zürich) and Martin Vingron (Free University of Berlin). In a fascinating talk, Bühlmann argued for the need of uncertainty quantification in an increasingly heterogeneous data world –examples of this are also appearing in other fields, e.g. in the study of autism disorders as Connie Kasari and Susan Murphy mentioned in “SMART Approaches to Combating Autism”- and for models to be the basis of the assignation of uncertainties – topic greatly covered by Andrew Gelman in “Living with uncertainty, yet still learning”-, while acknowledging that in the big data context, “confirmatory statistical inference is (even more) challenging”. Vingron followed by focusing on “Epigenomics as an example”, raising open questions to the audience on how to define causality in Biology where “most processes […] are feedback circular processes”, calling for models that are just complex enough so as to allow for mechanistic explanations, and for good definitions of null hypotheses.
In addition to the interesting points of the talk, I found its title particularly attractive in what it could be directly linked to a vibrant roundtable on “Genomics, Biostatistics and Bioinformatics” in the framework of the 2nd Biostatnet General Meeting, in which, as some of you might remember, the definitions of the terms Biostatistics and Bioinformatics were discussed. I wonder if the term “statistical Bioinformatics” would be indeed the solution to that dilemma? As a matter of fact, Bühlmann himself mentions at the start of his talk other options like Statistical Information Sciences, etc-
Michael Newton and David Schwartz from the University of Wisconsin-Madison also focused on the triumph of sequencing and “Millimeter-long DNA Molecules: Genomic Applications and Statistical Issues”. Breast cancer being one of the mentioned applications, this was followed by an introduction to “Statistics in Cancer Genomics” by Jason Carroll and Rory Stark (Cancer Research UK Cambridge Institute, University of Cambridge), particularly focusing on breast and prostate cancer and the process of targeting protein binding sites. The latter, as a computational biologist, mentioned challenges on the translation of Statistics for the biologists and vice versa, and identified ”reproducibility as (the) most critical statistical challenge” in the area – also on the topic of reproducibility, it is especially worth watching “Statistics, Big Data and the Reproducibility of Published Research” by Emmanuel Candes (Stanford University) -.
Another area increasingly gaining attention in parallel with technologies improvements is Neuroimaging. Thomas Nichols (University of Warwick) lead the audience in a trip through classical examples of its application (from Phineas Gage´s accident to observational studies of the structural changes in the hippocampi of taxi drivers, and the brain´s reaction to politics) up to current exciting times, with great “Opportunities in Functional Data Analysis” which Jane-Ling Wang (University of California) promoted in her talk.
Also in the area of Health Statistics, different approaches to dietary patterns modelling were shown in “The Analysis of Dietary Patterns and its Application to Dietary Surveillance and Epidemiology” by Raymond J. Carroll (Texas A&M University), and Sue Krebs-Smith (NCI), with challenges in the area being; finding dietary patterns over the life course – e.g. special waves of data would appear in periods like pregnancy and menarchy for women-, and incorporation of technology to the studies as a new form of data collection –e.g., pictures of food connected to databases-.
Again with a focus on the challenges posed by technology, three talks on environmental Statistics outlined an evolution over time of the field. Starting from current times, Marian Scott (University of Glasgow) in “Environment – Present and Future” stated in her talk that “natural variability is more and more apparent” because we are now able to visualise it. This idea going back to the heterogeneity claimed by Bühlmann. Despite the great amounts of data being available, and initially caused by technological improvements but ultimately due to a public demand – “people are wanting what it is happening now”-, the future of the field is still subject to the basic questions: “how and where to sample.” Especially thought-provoking were Scott´s words on “the importance of people in the environmental arena” and the need for effective communication: “we have to communicate: socio-economic, hard science…, it all needs to be there because it all matters, it applies to all the world…”
Amongst other aims of this environmental focus such as living sustainably – both in urban and rural environments-, climate is a major worry/fascination which is being targeted through successful collaborations of specialists in its study and statisticians. “Climate: Past to Future” by Gabi Hegerl (University of Edinburgh) and Douglas Nychka (NCAR) covered “Regional Climate – Past, Present and Future” and showed relevant examples of these successful collaborations.
Susan Waldron (University of Glasgow), under the captivating speech title of “A Birefringent Future”, highlighted the need to address challenges in the communication of “multiple layers of information” too, Statistics appearing as the solution for doing this effectively –e.g. by creating “accessible visualisation of data” and by incorporating additional environmental variation in controversial green energy studies such as whether wind turbines create a microclimate-. Mark Hansen (Columbia University Journalism School) seconded Waldron´s arguments and called for journalists to play an important role in the challenge: “data are changing systems of power in our world and journalists. […] have to be able to think both critically as well as creatively with data”, so as to “provide much needed perspective”. Examples of this role are; terms such as “computational Journalism” already being coined and data visualisation tools being put in place -as a fascinating example, the New York Times´ Cascade tool builds a very attractive representation of the information flow in Social Media-. David Spiegelhalter (Cambridge University) also dealt with the topic in the great talk “Statistics for an Informed Public”, providing relevant examples (further explanation on the red meat example can be found here as well).
To encourage “elicitation of experts opinions (to reach a) rational consensus” and “to spend more time with regulators” came up as other useful recommendations in the presentation “Statistics, Risk and Regulation” by Michel Dacorogna (SCOR) and Paul Embrechts (ETH Zürich).
In a highly connected world inmersed in a data revolution, privacy becomes another major issue. Statistical challenges arising from networked data –e.g. historical interaction information modelling- were addressed by Steve Fienberg (Carnegie Mellon University) and Cynthia Dwork (Microsoft Research), who argued that “statisticians of the future will approach data and privacy (and its loss) in a fundamentally algorithmic fashion”, in doing so explicitly answering the quote by Sweeney: “Computer science got us into this mess, can computer science get us out of it?”. Michael I. Jordan (University of California-Berkeley) in “On the Computation/Statistics Interface and “Big Data”” also referred to “bring(ing) algorithmic principles more fully into contact with statistical inference“.
I would like to make a final mention to one of the questions posed by the audience during the event. When Paul Embrechts was enquired about the differences between Econometrics and Statistics, a discussion followed on how crossfertilisation between fields has happened back and forth. As a direct consequence of this contact, fields rediscover issues from other areas. For instance, Credit Risk Analysis models were mentioned as being inferred from Medical Statistics or back in Peter Bühlmann´s talk, links were also found between Behavioural Economics and Genetics. These ideas, from my point of view, bring together the essence of Statistics, i.e. its application and interaction with multiple disciplines as the foundations of its success.
Many other fascinating topics were conveyed in the workshop but unfortunately, I cannot fit here mentions to all the talks. I am sure you will all agree it was a fantastic event.
 Natàlia Vilor-Tejedor holds a BSc in Mathematics and Applied Statistics from the Universitat Autònoma de Barcelona. Additionally, she holds a MSc in Omics Data Analysis from the Universitat de Vic where she also worked at the Bioinformatics and Medical Statistics Research group developing new methods for analyzing the individual evidence of SNPs in biological pathways and was involved in some GWAS data analyses. Currently, she is a PhD student in the Biomedicine programme at the Universitat Pompeu Fabra. At the same time, she is working at the Barcelona Biomedical Research Park (in the Centre for Research in Environmental Epidemiology) where she is working on new statistical methods for integrating different types of Omics, Neuroimaging and Clinical Data from the European BREATHE project. She received one of the two Biostatnet prizes for the best presentations at the JEDE III conference for her talk on “Efficient and Powerful Testing for Gene Set Analysis in Genome-Wide Association Studies”.
Natàlia Vilor-Tejedor holds a BSc in Mathematics and Applied Statistics from the Universitat Autònoma de Barcelona. Additionally, she holds a MSc in Omics Data Analysis from the Universitat de Vic where she also worked at the Bioinformatics and Medical Statistics Research group developing new methods for analyzing the individual evidence of SNPs in biological pathways and was involved in some GWAS data analyses. Currently, she is a PhD student in the Biomedicine programme at the Universitat Pompeu Fabra. At the same time, she is working at the Barcelona Biomedical Research Park (in the Centre for Research in Environmental Epidemiology) where she is working on new statistical methods for integrating different types of Omics, Neuroimaging and Clinical Data from the European BREATHE project. She received one of the two Biostatnet prizes for the best presentations at the JEDE III conference for her talk on “Efficient and Powerful Testing for Gene Set Analysis in Genome-Wide Association Studies”.