Isaac Subirana holds a MSc in Science and Statistical Techniques from the Universitat Politècnica de Catalunya (UPC) in 2005. Additionally, he was awarded his PhD in Statistics from the Universitat de Barcelona (UB) in 2014. Since 2007 he has been teaching statistics and mathematics at the faculty of Biology (UB) and Statistics (UPC) as associate professor. Since 2004 he has been working at REGICOR group (IMIM-Parc de Salut Mar) assessing statistical analyses in cardiovascular and genetic epidemiological studies. He has given several workshops of Shiny, at UPC Summer School, Servei d’Estadística Aplicada de la Universitat Autònoma de Bellaterra (SEA-UAB), at Basque Country University (EHU) and at the Institut Català d’Oncologia (ICO).
Isaac Subirana holds a MSc in Science and Statistical Techniques from the Universitat Politècnica de Catalunya (UPC) in 2005. Additionally, he was awarded his PhD in Statistics from the Universitat de Barcelona (UB) in 2014. Since 2007 he has been teaching statistics and mathematics at the faculty of Biology (UB) and Statistics (UPC) as associate professor. Since 2004 he has been working at REGICOR group (IMIM-Parc de Salut Mar) assessing statistical analyses in cardiovascular and genetic epidemiological studies. He has given several workshops of Shiny, at UPC Summer School, Servei d’Estadística Aplicada de la Universitat Autònoma de Bellaterra (SEA-UAB), at Basque Country University (EHU) and at the Institut Català d’Oncologia (ICO).
In the last decade, R has become one of the most used software both for statistical and data-analysis in general. On the one hand, R offers a great flexibility and power to perform any kind of analyses and computations. On the other, R has a very steep learning curve for beginners while other software such as SPSS are much more intuitive and easier to learn. By far, these point-and-click alternatives are the most commonly used by analysts who do not have the knowledge to manage R commands with confidence. It would appear that many physicians and researchers from other applied areas belong to this group of people who feel much more comfortable using this sort of software than writing commands.
The problem arises when a complex statistical analysis not implemented in SPSS-like software is required for a paper publication, e. g. spline models to assess dose-response effects. In such cases a researcher unfamiliar with R may enlist the help of a statistician to do the analysis. This prevents the researcher from performing data exploration or repeating the analyses by selecting groups of individuals, for instance, to create or confirm hypotheses. To overcome this situation, the statistician could provide the researcher with an R syntax indicating where to modify the code. However, this is not an optimal solution because the researcher would have to deal with an unfamiliar language and run a code that may return unintelligible error messages.
Some efforts have been done to bring R to less proficient users by building Graphical User Interfaces (GUIs). One of the most well-known examples is Rcmdr which is an R package to perform general statistical analyses. In addition, there are numerous Rcmdr plug-ins packages performing more specific analyses such as survival analyses, etc. Both, Rcmdr and their plug-ins, are built using tcltk tools (tcltk2 R package). By using tcltk2 it is possible to create and customize windows, buttons, lists, etc., but its syntax is not very intuitive, at least for R users. Other existing alternatives consist on using web program languages (HTML, Javascript and PHP) which also allow to plug-in R commands. The problem is that most R users are not familiar with HTML, Javascript or PHP, and building even a simple application may be too demanding.
In 2012, a new R package called Shiny was submitted to R CRAN repository (see Pilar’s post on the topic here). It was created and is maintained by Joe Cheng and collaborators from the RStudio team. Unlike tcltk2, Shiny is much more intuitive and simple. Shiny wraps HTML and Javascript using exclusively R instructions to create web based GUIs which can be opened from any internet explorer (such as Chrome, Firefox, etc.). Therefore, Shiny takes all the power of HTML, Javascript and R without having to know anything of the first two. From a usability point of view, the main advantage of creating GUIs applications with Shiny is that they can be called from any device (see this page for more details). On the Shiny website there are a lot of examples and a very extensive list of tutorials and article. I would strongly recommend that you visit it before starting to create an application with Shiny.
Since Shiny was first presented in useR 2013 conference in Albacete its popularity has grown exponentially. More and more R packages incorporates its GUI built with Shiny; compareGroups to build descriptive tables, MAVIS for meta-analyses, Factoshiny which is a Shiny-web GUI of FactoMineR package for factor analyses, or GOexpress for genomic data analyses are some examples. Even a section in R-bloggers has been created exclusively for Shiny topics (see this website). And specific Shiny conferences are taking place (see this website).
By default, Shiny applications may look too minimalistic, and sometimes, their functionality could be seen as limited. To improve Shiny applications aspect and functionality, there are some R packages available on CRAN. Of special interest are shinythemes which incorporates a list of CSS templates, shinyBS to build modals or shinyjs which wraps Javascript code.
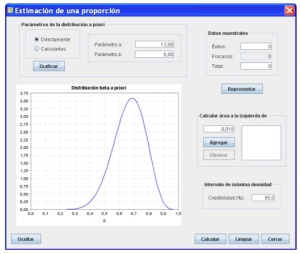
I started using Shiny to create a GUI for compareGroups, an R package to build descriptive tables for which I am the maintainer. We found the necessity to open the compareGroups package to SPSS-like software users not familiar with R. To do so, it was necessary to create an intuitive GUI which could be used remotely without having to install R, upload your data in different formats (specially, SPSS and Excel), select variables, and other options with drop-down lists and buttons. You can take a look at the compareGroups project website for further information. Aside from developing the compareGroups GUI, during these last three years I have also been designing other Shiny applications, ranging from performing models (website) to teaching statistics in the university (website).
In conclusion, Shiny is a great tool to be used by R-users who are not familiar with HTML, Javascript or PHP to create very flexible and powerful web-based applications.
If you know how to do something in R, you can make other non-R users do it by themselves too!